Annex 3: Guideline development group members
Table A.3.1. Guideline development group members: “Targeted nextgeneration sequencing” 2–5 May 2023
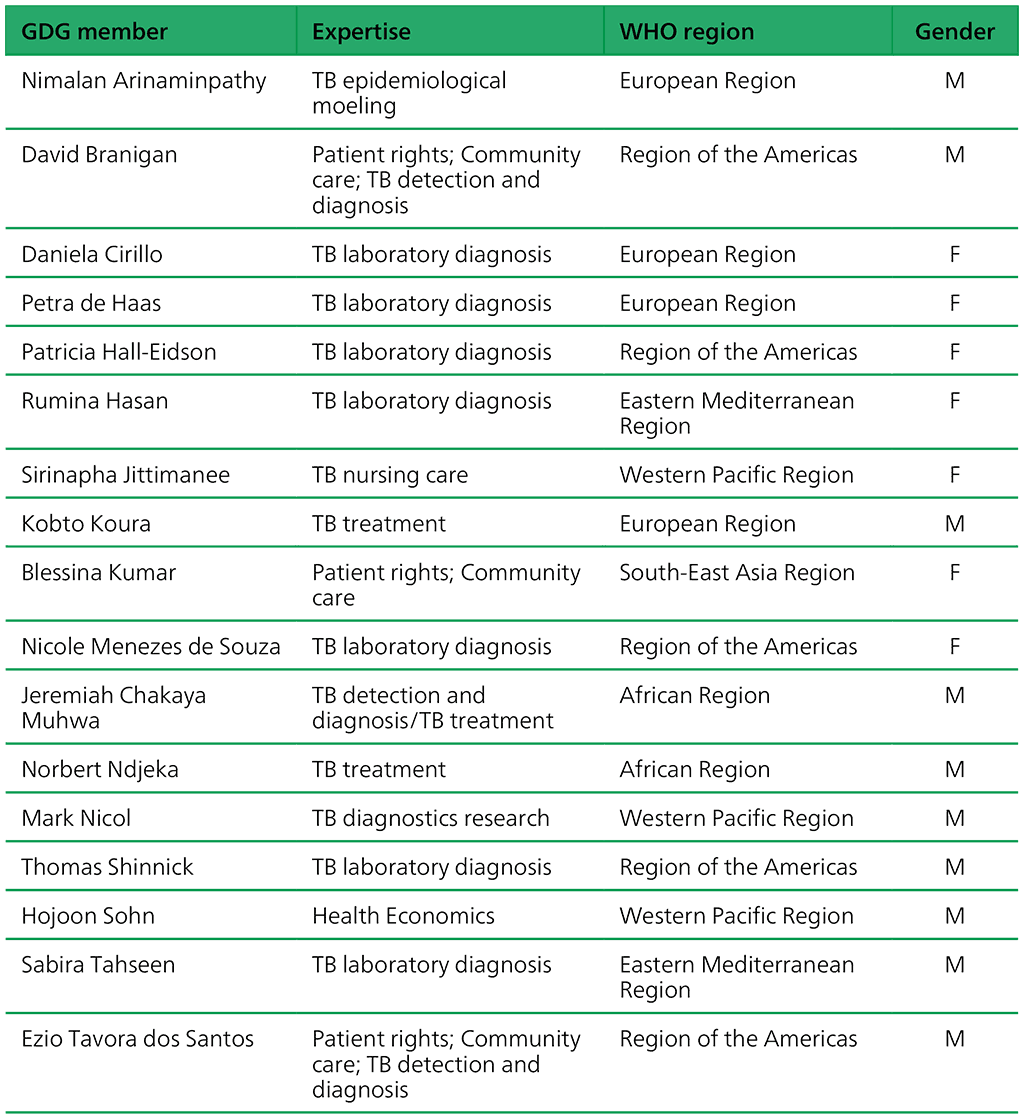

Table A.3.1. Guideline development group members: “Targeted nextgeneration sequencing” 2–5 May 2023


In patients with MDR/RR-TB, a positive SL-LPA result for fluoroquinolone resistance (as a class) or SLID resistance (as a group) can be treated with confidence. The diagnostic accuracy of SL-LPA is similar when performed directly on sputum specimens or indirectly on cultured isolates of M. tuberculosis.
Targeted NGS technology couples amplification of selected genes with NGS technology to detect resistance to many drugs with a single test. Also, since targeted NGS can interrogate entire genes to identify specific mutations associated with resistance, tests based on this technology may be more accurate than existing WRDs. In addition, new tests based on NGS can detect resistance to new and repurposed drugs that are not currently included in any other molecular assays.
Bruker-Hain Diagnostics has two real-time nucleic acid amplification tests (NAATs), the FluoroType MTB to detect Mycobacterium tuberculosis complex (MTBC) and the FluoroType MTBDR, to detect MTBC, and resistance to rifampicin (RIF) and isoniazid (INH) in tuberculosis (TB). The MTB test (VER 1.0) targets the IS6110 DNA insertion element for MTBC detection, while the MTBDR test (VER 2.0) targets the rpoB gene for detection of MTBC and RIF resistance, and the inhA promoter and katG gene for detection of INH resistance.
The Xpert MTB/XDR detects Mycobacterium tuberculosis complex (MTBC) DNA and genomic mutations associated with resistance to isoniazid (INH), fluoroquinolones (FQs), ethionamide (ETH) and second-line injectable drugs (amikacin [AMK], kanamycin and capreomycin) in a single cartridge. Xpert MTB/XDR tests are run on Cepheid's GeneXpert instruments, using 10-colour modules that differ from the 6-colour modules traditionally used for Xpert MTB/RIF and Xpert MTB/RIF Ultra (Xpert Ultra) testing.
Nipro (Osaka, Japan) developed Genoscholar PZA-TB, a reverse hybridization-based technology for detection of pyrazinamide (PZA) resistance in tuberculosis (TB) (1, 2). Compared with MTBDRplus and MTBDRs/ LPA, the Genoscholar PZA-TB line-probe assay (LPA) does not include specific mutant probes, because resistance mutations are widespread across the entire pncA gene with no predominant mutations.
Roche Molecular Systems, Inc. (RMS, Roche) has two nucleic acid amplification tests (NAATs), the cobas MTB and cobas MTB-RIF/INH tests, to detect Mycobacterium tuberculosis complex (MTBC) and drug resistance (rifampicin [RIF] and isoniazid [INH]), respectively (1,2) in tuberculosis (TB). The MTB assay detects both 16S rRNA and esx genes as target genes for MTBC detection.
Becton Dickinson (BD) has a multiplexed real-time polymerase chain reaction (PCR) nucleic acid amplification tests (NAAT) (BD MAX MDR-TB) for the detection of Mycobacterium tuberculosis complex (MTBC) and resistance to both rifampicin (RIF) and isoniazid (INH) in tuberculosis (TB). For MTBC detection, this test targets the multicopy genomic elements IS6110 and IS1081, as well as a single copy genomic target.